-
-
Notifications
You must be signed in to change notification settings - Fork 36
Commit
This commit does not belong to any branch on this repository, and may belong to a fork outside of the repository.
challenges: add nucleotide count exercism
- Loading branch information
1 parent
72bfe72
commit 75a4e00
Showing
2 changed files
with
374 additions
and
0 deletions.
There are no files selected for viewing
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,373 @@ | ||
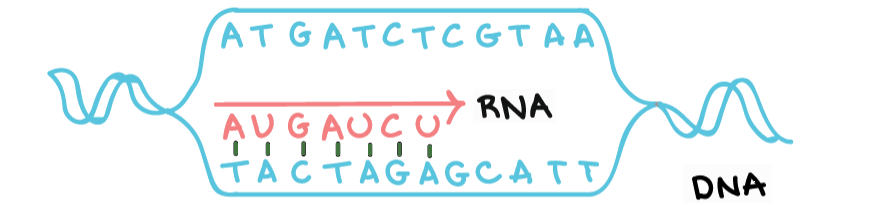
| # Nucleotide Count | ||
|
|
||
|  | ||
|
|
||
| [:globe_with_meridians: Clojure Track: Nucleotide Count](https://exercism.org/tracks/clojure/exercises/nucleotide-count){target=_blank .md-button} | ||
|
|
||
| Given a string representing a DNA sequence, count how many of each nucleotide is present. | ||
|
|
||
| If the string contains characters other than A, C, G, or T then an error should be throw. | ||
|
|
||
| Represent a DNA sequence as an ordered collection of nucleotides, e.g. a string of characters such as "ATTACG". | ||
|
|
||
|
|
||
| ```shell | ||
| "GATTACA" -> 'A': 3, 'C': 1, 'G': 1, 'T': 2 | ||
| "INVALID" -> error | ||
| ``` | ||
|
|
||
| !!! INFO "DNA Nucleotide names" | ||
| `A` is Adenine, `C` is Cytosine, `G` is Guanine and `T` is Thymine | ||
|
|
||
|
|
||
| !!! HINT "Code for this solution on GitHub" | ||
| [practicalli/exercism-clojure-guides](https://github.com/practicalli/exercism-clojure-guides/) contains the design journal and solution to this exercise and many others. | ||
|
|
||
|
|
||
| ## Create the project | ||
|
|
||
| Download the Nucleotide Count exercise using the exercism CLI tool | ||
|
|
||
| !!! NOTE "" | ||
| ```bash | ||
| exercism download --exercise=nucleotide-count --track=clojure | ||
| ``` | ||
|
|
||
| !!! HINT "Use the REPL workflow to explore solutions locally" | ||
| Open the project in a [Clojure aware editor](/clojure/clojure-editors) and [start a REPL](/clojure/coding-challenges/exercism/#repl-workflow), using a rich comment form to experiment with code to solve the challenge. | ||
|
|
||
|
|
||
| ## Starting point | ||
|
|
||
| Unit test code calls functions from the `src` tree which must exist with the correct argument signature for the unit test code to compile successfully. | ||
|
|
||
| Reviewing each assertion in the unit test code identifies the function definitions required. | ||
|
|
||
| ??? EXAMPLE "Exercism Unit Tests" | ||
| ```clojure | ||
| (ns nucleotide-count-test | ||
| (:require [clojure.test :refer [deftest is]] | ||
| nucleotide-count)) | ||
|
|
||
| (deftest empty-dna-strand-has-no-adenosine | ||
| (is (= 0 (nucleotide-count/count-of-nucleotide-in-strand \A, "")))) | ||
|
|
||
| (deftest empty-dna-strand-has-no-nucleotides | ||
| (is (= {\A 0, \T 0, \C 0, \G 0} | ||
| (nucleotide-count/nucleotide-counts "")))) | ||
|
|
||
| (deftest repetitive-cytidine-gets-counted | ||
| (is (= 5 (nucleotide-count/count-of-nucleotide-in-strand \C "CCCCC")))) | ||
|
|
||
| (deftest repetitive-sequence-has-only-guanosine | ||
| (is (= {\A 0, \T 0, \C 0, \G 8} | ||
| (nucleotide-count/nucleotide-counts "GGGGGGGG")))) | ||
|
|
||
| (deftest counts-only-thymidine | ||
| (is (= 1 (nucleotide-count/count-of-nucleotide-in-strand \T "GGGGGTAACCCGG")))) | ||
|
|
||
| (deftest validates-nucleotides | ||
| (is (thrown? Throwable (nucleotide-count/count-of-nucleotide-in-strand \X "GACT")))) | ||
|
|
||
| (deftest counts-all-nucleotides | ||
| (let [s "AGCTTTTCATTCTGACTGCAACGGGCAATATGTCTCTGTGTGGATTAAAAAAAGAGTGTCTGATAGCAGC"] | ||
| (is (= {\A 20, \T 21, \G 17, \C 12} | ||
| (nucleotide-count/nucleotide-counts s))))) | ||
| ``` | ||
|
|
||
|
|
||
| !!! EXAMPLE "Function definitions required to compile unit test code" | ||
| ```clojure title="src/nucleotide_count.clj" | ||
| (ns nucleotide-count) | ||
|
|
||
| (defn count-of-nucleotide-in-strand | ||
| "Count how many of a given nucleotide is in a strand" | ||
| [nucleotide strand]) | ||
|
|
||
| (defn nucleotide-counts | ||
| "Count all nucleotide in a strand" | ||
| [strand]) | ||
| ``` | ||
|
|
||
|
|
||
| ## Making the tests pass | ||
|
|
||
| Select one assertion from the unit tests and write code to make the test pass. | ||
|
|
||
| Experiment with solutions in the `comment` form and add the chosen approach to the respective function definition. | ||
|
|
||
|
|
||
| ## Counting nucleotides | ||
|
|
||
| Use test data from the unit test code, e.g. `"GGGGGTAACCCGG"` | ||
|
|
||
| How often does a nucleotide appear | ||
|
|
||
| !!! EXAMPLE | ||
| ```clojure | ||
| (map | ||
| #(if (= % \A) 1 0) | ||
| "GGGGGTAACCCGG") | ||
| ``` | ||
|
|
||
|
|
||
| Add the result to get the total count | ||
|
|
||
| !!! EXAMPLE | ||
| ```clojure | ||
| (count | ||
| (map | ||
| #(if (= % \A) 1 0) | ||
| "GGGGGTAACCCGG")) | ||
| ``` | ||
|
|
||
| Is there a more elegant way? | ||
|
|
||
| When only the matching nucleotide is in the strand, then all the elements of the strand can be counted. | ||
|
|
||
| `filter` the DNA strand with a predicate function (returns true/false) that returns only the matching nucleotide. | ||
|
|
||
| !!! EXAMPLE | ||
| ```clojure | ||
| (filter #(= % \A) valid-nucleotides)) | ||
| ``` | ||
|
|
||
| ;; Count the elements in the returned sequence for the total | ||
|
|
||
|
|
||
| !!! EXAMPLE | ||
| ```clojure | ||
| (count | ||
| (filter #(= % \A) valid-nucleotides)) | ||
| ``` | ||
|
|
||
|
|
||
| Add this code into the starting function | ||
|
|
||
|
|
||
| ### Run unit tests | ||
|
|
||
| Run the unit tests to see if they pass. x should pass, x should fail. | ||
|
|
||
|
|
||
| ### Nucleotide occurances | ||
|
|
||
| Count the occurances | ||
|
|
||
| "GGGGGTAACCCGG" | ||
|
|
||
|
|
||
| ```clojure | ||
| (count | ||
| (filter (fn [nucleotide] (= nucleotide \A)) | ||
| "GGGGGTAACCCGG")) | ||
| ``` | ||
|
|
||
|
|
||
| Define the data | ||
|
|
||
| ```clojure | ||
| (def valid-nucleotides | ||
| "Characters representing valid nucleotides" | ||
| [\A \C \G \T]) | ||
| ``` | ||
|
|
||
| Exception handling required | ||
|
|
||
| ```clojure | ||
| (throw (Throwable.)) if nucleotide is \X | ||
| ``` | ||
|
|
||
| Or use a predicate with some (some element? in the sequence) | ||
|
|
||
| ```clojure | ||
| (some #(= \G %) valid-nucleotides) | ||
|
|
||
| (some #{\G} valid-nucleotides) | ||
| ``` | ||
|
|
||
| ```clojure | ||
| (defn count-of-nucleotide-in-strand | ||
| [nucleotide strand] | ||
| (if (some #(= nucleotide %) valid-nucleotides) | ||
| (count | ||
| (filter #(= nucleotide %) | ||
| strand)) | ||
| (throw (Throwable.)))) | ||
|
|
||
| (count-of-nucleotide-in-strand \T "GGGGGTAACCCGG") | ||
| ``` | ||
|
|
||
|
|
||
| Design the second function | ||
|
|
||
| How often does a nucleotide appear | ||
|
|
||
| ```clojure | ||
| (map | ||
| #(if (= % \A) 1 0) | ||
| valid-nucleotides) | ||
| ``` | ||
|
|
||
| Add the result to get the total count | ||
|
|
||
| Is there a more elegant way? | ||
|
|
||
| ```clojure | ||
| (filter #(= % \A) valid-nucleotides) | ||
| ``` | ||
|
|
||
| Count the elements in the returned sequence for the total | ||
|
|
||
| Design the second function | ||
|
|
||
| How often does a nucleotide appear | ||
|
|
||
| NOTE: zero must be returned when there are no appearences | ||
|
|
||
| Return value always in the form | ||
|
|
||
| ```clojure | ||
| {\A 20, \T 21, \G 17, \C 12} | ||
| ``` | ||
|
|
||
| ### Hammock time... | ||
|
|
||
| - How often does something appear, | ||
| - how frequenct is it? | ||
| - Is there a clojure standard library for that (approx 700 functions), review <https://clojure-docs.org/> | ||
|
|
||
|
|
||
| ```clojure | ||
| (frequencies "GGGGGAACCCGG") | ||
| ``` | ||
|
|
||
| If there are missing nucleotides then there is no answer | ||
|
|
||
| What if there is a starting point | ||
|
|
||
| ```clojure | ||
| {\A 0 \C 0 \G 0 \T 0} | ||
| ``` | ||
|
|
||
| ;; Then merge the result of frequencies | ||
|
|
||
| ```clojure | ||
| (merge {\A 0 \C 0 \G 0 \T 0} | ||
| (frequencies "GGGGGAACCCGG")) | ||
| ``` | ||
|
|
||
| Update the function definition and run tests | ||
|
|
||
|
|
||
| ## Solutions | ||
|
|
||
| There are many ways to solve a challenge and there is value trying different approaches to help learn more about the Clojure language. | ||
|
|
||
| The following solution includes `filter` and `frequencies` functions which are commonly used functions from the Clojure standard library. | ||
|
|
||
| !!! EXAMPLE "Example Solution" | ||
| ```clojure title="src/nucleotide_count.clj" | ||
| (ns nucleotide-count) | ||
|
|
||
| (def valid-nucleotides | ||
| "Characters representing valid nucleotides" | ||
| [\A \C \G \T]) | ||
|
|
||
| (defn count-of-nucleotide-in-strand | ||
| [nucleotide strand] | ||
| (if (some #(= nucleotide %) valid-nucleotides) | ||
| (count | ||
| (filter #(= nucleotide %) | ||
| strand)) | ||
| (throw (Throwable.)))) | ||
|
|
||
| (defn nucleotide-counts | ||
| "Count all nucleotide in a strand" | ||
| [strand] | ||
| (merge {\A 0 \C 0 \G 0 \T 0} | ||
| (frequencies "GGGGGAACCCGG"))) | ||
| ``` | ||
|
|
||
|
|
||
|
|
||
|
|
||
|
|
||
|
|
||
| <!-- | ||
| ;; --------------------------------------------------------- | ||
| ;; Solution | ||
| #_(defn count-of-nucleotide-in-strand [nucleotide strand] | ||
| (if (= \X nucleotide) | ||
| (throw (Throwable.)) | ||
| (count (filter #(= nucleotide %) strand)))) | ||
| #_(defn nucleotide-counts [strand] | ||
| (apply merge | ||
| (for [nucleotide strand] | ||
| {nucleotide | ||
| (count-of-nucleotide-in-strand nucleotide strand)}))) | ||
| #_(defn count-of-nucleotide-in-strand [nucleotide strand] | ||
| (if (some (set nucleotide) valid-nucleotides) | ||
| (count (filter #(= nucleotide %) strand)) | ||
| (throw (Throwable.))))) | ||
| ;; End of Solution | ||
| ;; --------------------------------------------------------- | ||
| (comment) | ||
| #_{:clj-kondo/ignore [:redefined-var]} | ||
| #_() ; End of rich comment | ||
| ;; (defn count-of-nucleotide-in-strand [nucleotide strand] | ||
| ;; (get (frequencies strand) nucleotide (throw (Throwable.)))) | ||
| ;; add a guard condition if passed an empty strand | ||
| ;; returning zero | ||
| ;; (defn count-of-nucleotide-in-strand [nucleotide strand] | ||
| ;; (if (empty? strand) | ||
| ;; 0 | ||
| ;; (get (frequencies strand) nucleotide (throw (Throwable.))))) | ||
| ;; (count-of-nucleotide-in-strand \T "GGGGGTAACCCGG") | ||
| ;; using a function as the not-found | ||
| ;; (get {:a 1 :b 2} :b (throw (Throwable.))) | ||
| ;; unfortunately the throw expression is evaluated before the get | ||
| ;; so we always get a Throwable exception even if the key is in the hash-map. | ||
| ;; We can fix that with an or | ||
| ;; (defn count-of-nucleotide-in-strand [nucleotide strand] | ||
| ;; (if (empty? strand) | ||
| ;; 0 | ||
| ;; (or (get (frequencies strand) nucleotide) | ||
| ;; (throw (Throwable.))))) | ||
| ;; | ||
| ;; (defn nucleotide-counts [strand] | ||
| ;; (frequencies strand)) | ||
| ;; | ||
| ;; fails the empty case, | ||
| ;; but we can add an empty result as the default value | ||
| ;; and merge it with the resulting frequencies | ||
| ;; (defn nucleotide-counts [strand] | ||
| ;; (merge {\A 0 \C 0 \G 0 \T 0} (frequencies strand))) | ||
| --> |
This file contains bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters